Proprietary Drug Discovery Platform - System(PDPS)
Overview of PDPS
At PeptiDream, we use our proprietary drug discovery platform, PDPS (Peptide Discovery Platform System1) to identify and develop innovative drugs based on macrocyclic peptides for our internal research programs and with our collaboration partners. PDPS enables the generation of highly diverse libraries of macrocyclic peptides which can be screened against almost any biological target of interest to discover hit compounds rapidly (~1 month) with a high success rate (>95%).
1) PeptiDream acquired the exclusive license to the PDPS patent portfolio from The University of Tokyo, and further refined the technology and developed internal expertise to become the leading player in peptide drug discovery.
Outline of PDPS
In brief, PDPS has 3 key features:
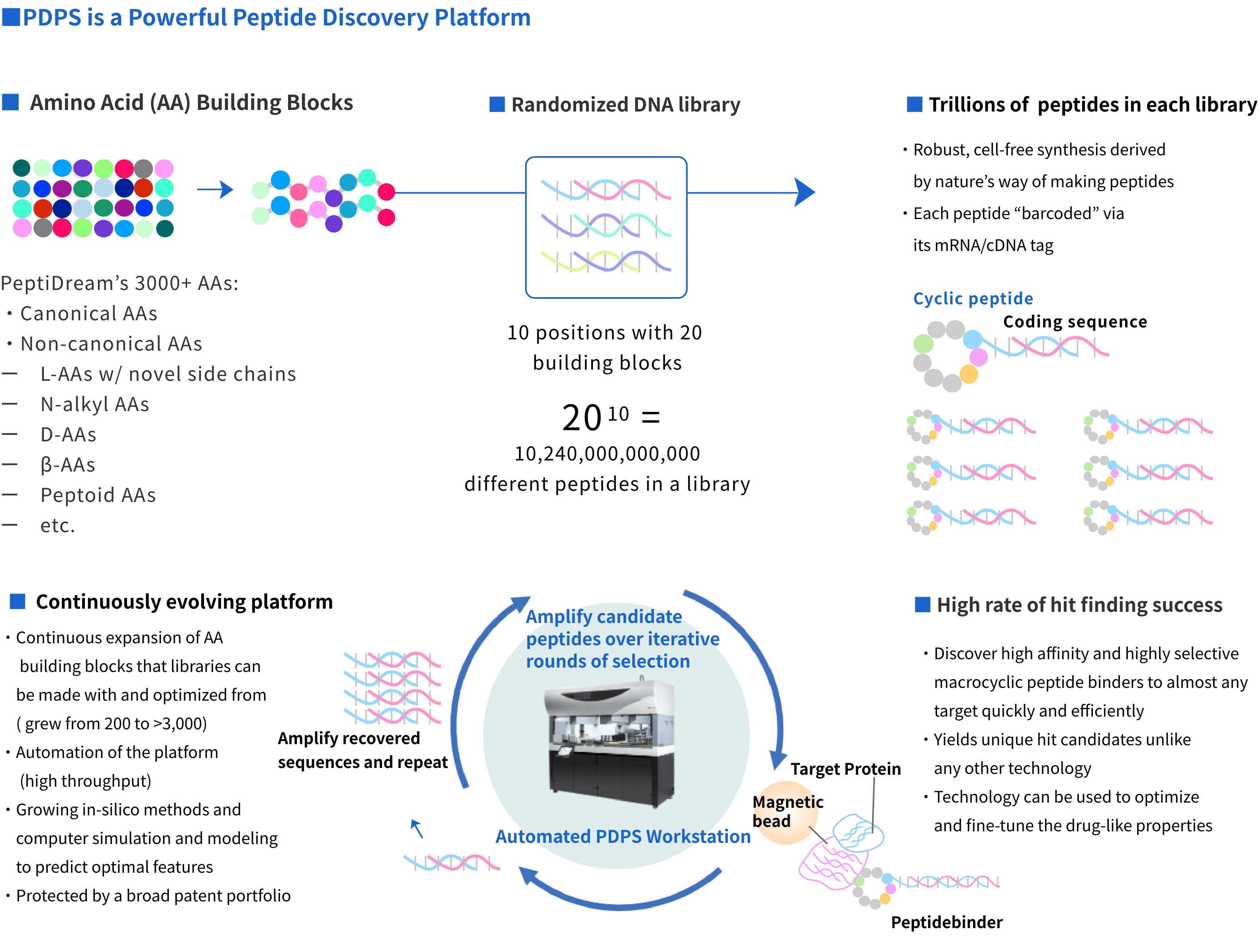
1) Access to trillions of macrocyclic peptides in a library
Our platform can generate trillions of macrocyclic peptides with limitless chemical diversity unrivaled by any
other hit finding technology. The peptides are synthesized with high efficiency in a cell-free,
transcription/translation system using randomized DNA libraries and canonical and/or non-canonical amino acid
building blocks. All peptides are “barcoded” with their respective mRNA/cDNA tags.
2) High success rate of finding hits
The peptide libraries are screened against the biological target of interest and counter-screened against
other targets to select for peptides with high affinity and selectivity for the desired target (“hit
peptides”). The “barcodes” attached to the peptides can be used to amplify the hit peptides over iterative
rounds of selection and allows for rapid identification of the amino acid sequences of the hit peptides. The
platform can also be used for optimization and assessment of the hits, which can significantly accelerate the
drug discovery process.
3) Constant evolution
At PeptiDream, we are constantly refining and improving the capabilities of PDPS. We have expanded the
chemical diversity of the amino acid building blocks that can be accommodated into the platform and developed
an automated system which increases the throughput and reduces variability of results. We are also developing
in silico methods to facilitate informed design of our libraries and strive to be at the forefront of
innovation in drug discovery.

